Search Details
| UAMH Number: | 10320 |
|---|---|
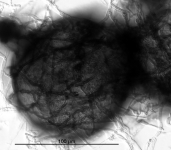
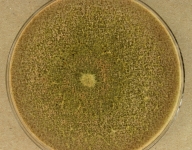
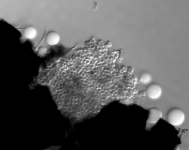
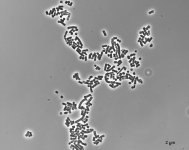
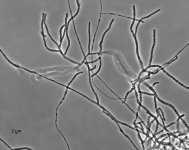
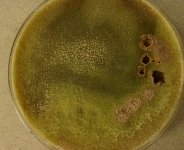
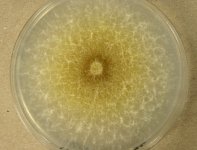
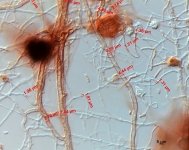
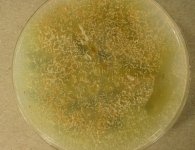
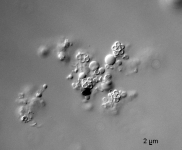
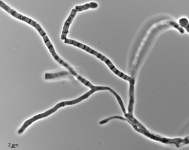
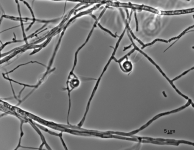
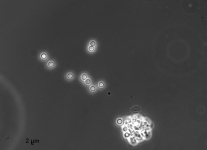
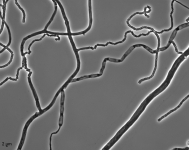
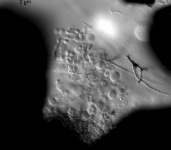
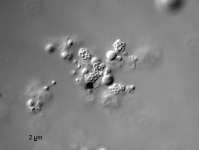
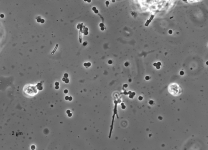
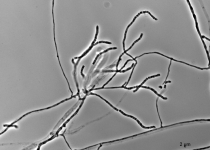
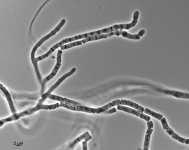
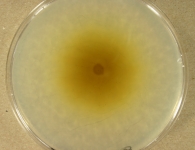
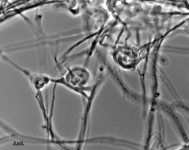
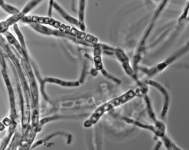
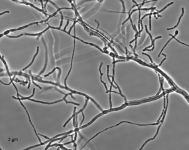
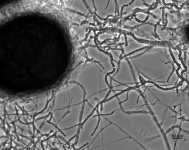
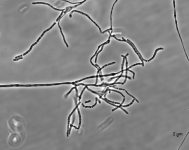
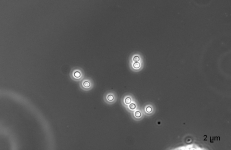
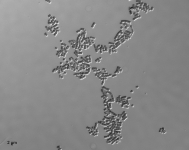
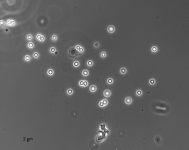
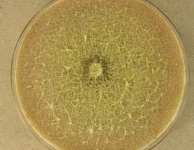
| Species Name: | Xylogone ganodermophthora |
| Type: | Xylogone ganodermophthora |
| Synonyms: | |
| Taxonomy: | FUNGI Ascomycota, Leotiomycetes, Helotiales, Hyaloscyphaceae |
| Strain History: | Kang, H-J (H55) -> UAMH |
| Substrate: | oak wood logs used for mushroom cultivation (Ganoderma lucidum) | Location: | SOUTH KOREA Gyeonggi Province (GEO: 37.414,127.518) |
| Isolator: | H-J Kang |
| Isolation Date: | 2016-01-10 |
| Date Received: | 2003-07-08 |
| Characters: | BIODETERIOGEN/ BIODEGRADATION Yellow rot pathogen of cultivated Ganoderma lucidum in Korea - // MOLECULAR SYSTEMATICS Classification of yellow rot pathogen of cultivated Ganoderma lucidum - Kang HJ, Sigler L, Lee J, Gibas CFC, Yun SH, Lee Y, Mycologia 102:1167-1184, 2010 // PIGMENT yellow - (Click for publications citing UAMH 10320) |
| Compounds: | |
| Cross Reference: | CBS 127676 |
| Collections: | Living Strains; Dried Herbarium Material |
| Pathogenic Potential: | Human: no | Animal: no | Plant: no |
| Biosafety Risk Group: | RG1 (check the PHAC ePATHogen Risk Group Database for updates) |
| Regulatory Requirements: | Canadian requesters must provide PHAC Pathogen and Toxin License Number (see: https://www.canada.ca/en/public-health/services/laboratory-biosafety-biosecurity/licensing-program.html) prior to shipment. International requesters must provide all legally required importation documentation prior to shipment. Plant pathogenicity status may be verified by using the USDA Agricultural Research Service (ARS) Fungal Database |
| MycoBank ID: | 515401 |
| Sequences: | >UAMH10320_GQ290121_RNApol2 TTAACACAGGATGTCTACAAGTACTTACAGCGATGTGTTGAAAATAACCGTGAGTTCAATTTGACACTTGGAGTAAAATCCACAACATTGACCAACGGTCTTAAATACTCATTGGCTACTGGAAATTGGGGAGATCAAAAGAAGGCTGCAAGCTCGACAGCTGGTGTGTCTCAGGTGCTCAACAGATATACATTTGCCTCTACGCTTTCTCATTTGCGGCGAACCAATACACCTATTGGTCGTGATGGAAAGATTGCTAAACCTCGGCAGCTTCATAACACTCATTGGGGTCTGGTTTGTCCCGCCGAGACTCCAGAAGGTCAAGCTTGTGGTCTCGTCAAGAATCTCGCTCTCATGTGTTATGTTACTGTCGGTACTCCTAGTGACCCCATCGTTGAGTTCATGATTCAACGAAATATGGAAGTGCTCGAGGAATACGACCCAGTCAGATCACCAAATATGACCAAGGTCTTCGTCAATGGTGTTTGGGTAGGAGTTCATCGCGAACCCGCTCATCTCGTTAGCACCGTGCAACATCTCCGACGTTCTCATTTGATCTCACATGAAGTATCTCTAATTAGAGATATTCGTGACCGGGAATTCAAGATCTTCACAGATGCGGGCAGAGTCTGCAGACCATTGTTCGTTATTGATAATGATGTTGATAGCGCGAACAAAGGTAACTTGGTGCTCAACAAAGACCATATTCGAAGGCTAGAGGAAGATCAAACAATGCCTGCAAACATGGATATGGAACAGCGAAAGGAAGCAGGTTATTTCGGTTTCCAAGGTTTAATCAATGAGGGTGTGGTTGAATATGTAGACGCCGAAGAAGAGGAAACGATTATGATAGTGATGACTCCAGAAGATCTAGATATTTCTCGACAACTACAAGCCGGATATCAGATCAGACCTGATGAGAGTGGAGATATGAACAAACGTGTTAAGGCTCCAATGAACCCAACGGCGCATATCTGGACGCATTGTGAAATCCACCCAAGTATGATTCTGGGTATTTGTGCCAGTATCATTCCATTCCCGGATCATAATCAGNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNGGAGTGTTCCTTACTAATTTCGATCAACGTATGGATACCATGGCCAATATCTTGTACTATCCTCAGAAACCACTTGCTACCACACGTTCTATGGAATTCTTGAAGTTCAGAGAGTTGCCAGCAGGGCAGAATGCAATCGTCGCTATCGCATGTTACTCCGGTTACAATCAAGAAGATTCCGTTATTATGAATCAGAGTAGTATCGATCGGGGCCTTTTCCGAAGTCTCTTCTACAGAGCTTATACTGATCAAGAGAAGCGTATTGGAATGAATGTTGTTGAGCAGTTTGAGAAACCATTCAGATCAGACACCTTGAAGCTGAAACACGGTACTTATGACAAACTTGATGATGATGGTATCATTGCTCCAGGTGCCCGTGTCTCTGGAGAGGATATTATTATTGGAAAGACGGCACCCATTGCTCCTGACTCAGAAGAGCTTGGTCAGCGCACCAAGGCCCACGTCAAACGTGATGCCTCAACACCTCTTAGAAGCACTGAAAACGGTATTGTGGACCAGGTTCTTATCACTACGAATGCAGAGGGTCTTCGATTCGTTAAAGTTCGTATGCGAACTACGAAGATTCCACAGATTGGTGATAAATTTGCTTCCCGTCACGGACAAAAGGGTACCATTGGTATTACTTACCGACAAGAGGATATGCCATTTACTAGACAGGGTATCGTTCCGGATTTGATCATCAATCCTCATGCTATTCCATCTCGTATGACAATTGCTCACTTGATTGAATGTCAATTGAGCAAGGTCTCGACTCTTCGTGGTCTAGAGGGTGATGCGACACCTTTCACTGAAGTCACCGTTGATTCTGTTTCAAAACTTCTTCGTGCTCATGGCT >UAMH10320_GQ280399_SSU CNCAGATTAGCCATGCATGTCTAAGTATAAGCAACTATACTGTGAAACTGCGAATGGCTCATTAAATCAGTTATCGTTTATTTGATAGTACCTTACTACTTGGATAACCGTGGTAATTCTAGAGCTAATACATGCTAAAAACCTCGACTTCGGAAGGGGTGTATTTATTAGATAAAAAACCAATGCCCTTCGGGGCTCCTTGGTGATTCATAATAACTTAACGAATCGCATGGCCTTGTGCCGGCGATGGTTCATTCAAATTTCTGCCCTATCAACTTTCGATGGTAGGATAGTGGCCTACCATGGTTTCAACGGGTAACGGGGAATTAGGGTTCTATTCCGGAGAAGGAGCCTGAGACACGGCTCCTACATCCAAGGAAGGCAGCAGGCGCGCAAATTACCCAATCCCGACACGGGGAGGTAGTGACAATAAATACTGATACAGGGCTCTTTTGGGTCTTGTAATTGGAATGAGTACAATTTAAATCCCTTTAACGAGGAACAATTGGAGGGCAAGTCTGGTGCCAGCAGCCGCGGTAATTCCAGCTCCAATAGCGTATATTAAAGTTGTTGCAGTTAAAAAGCTCGTAGTTGAACCTTGGGCCTGGCTGGCCGGTCCGCCTCACCGCGTGCACTGGTCCGGCCGGGCCTTTCCTTCTGGGGAGCCGCATGCCCTTCACTGGGTGTGTCGGGGAACCAGGACTTTTACTTTGAAAAAATTAGAGTGTTCAAAGCAGGCCTATGCTCGAATACATTAGCATGGAATAATAGAATAGGACGTGTGGTTCTATTTTGTTGGTTTCTAGGACCGCCGTAATGATTAATAGGGATAGTCGGGGGCATCAGTATTCAATTGTCAGAGGTGAAATTCTTGGATTTATTGAAGACTAACTACTGCGAAAGCATTTGCCAAGGATGTTTTCATTAATCAGTGAACGAAAGTTAGGGGATCGAAGACGATCAGATACCGTCGTAGTCTTAACCATAAACTATGCCGACTAGGGATCGGGCGATGTTACTTTTTTGACTCGCTCGGCACCTTACGAGAAATCAAAGTCTTTGGGTTCTGGGGGGAGTATGGTCGCAAGGCTGAAACTTAAAGAAATTGACGGAAGGGCACCACAATGGAGTGGAGCCTGCGGCTTAATTTGACTCAACACGGGGAAACTCACCAGGTCCAGACACAATAAGGATTGACAGATTGAGAGCTCTTTCTTGATTTTGTGGGTGGTGGTGCATGGCCGTTCTTAGTTGGTGGAGTGATTTGTCTGCTTAATTGCGATAACGAACGAGACCTTNACCTACTAAATAGCCAGGCTAGCTTTGGCTGGTCGCCGGCTTCTTAGAGGGACTATCGGCTCAAGCCGATGGAAGTTTGAGGCAATAACAGGTTAACTTCACAGGCCTGTAAAAGCAGGTCTCAGACTTTCAGTGGGGAATGCTGGATAACTGCTAGTACACCATCTAATCGCTGTGGGGCGAGTGCCCCCATTATGAGGCAGCGACCGTTCAGGTGATGGGTGCGAGACAACCTGGTACAGGGGACGCCAGTCCGTCACTCGGTGATGGGCGGGTCAATCCTGTGGCGAGATAGGTAACGACTATCCGTCGCAACGCACGCTAAGGTGTCGGTCTACTGGATATCCGGTAGGCTTAAGGTACGTGCTATCCCCCGTGTAAGCGGGCCTCGAGAAATAGGACTCATAAGCCGAAGTCTCGAGGGATGCAGATTTAGGAGTTCTGTGAAATCACTGATCTGCATTGGGGTTGTTACTTTTGAGATTTTTCAGATAATTGGATTTCATTGGGAATCACGCGACAGCAGTCGCGGTGAACTCGATTTCTCTCAGAATACATGGTGGGGTGCAGGTGCATTCATGCAGCCGCAAGAGAATCCCAAACAATGAATCCAATTCGAAGAAGTTNTCAAAGGTGACAATGAAATGCTGTGATGCCCTTAGATGTTCTGGGCCGCACGCGCGCTACACTGACAGAGCCAACGAGTTCATCACCTTGGTCGAAAGGCCTGGGTAATCTTGTTAAACTCTGTCGTGCTGGGGATAGAGCATTGCAATTATTGCTCTTCAACGAGGAATTCCTAGTAAGCGCAAGTCATCAGCTTGCGTTGAATACGTCCCTGCCCTTTGTACACACCGCCCGTCGCTACTACCGATTGAATGGCTCAGTGAGGCTTTCGGACTGGCCTAGGAAGAGTGGCAACACTCATCCAGGCCGGAAAGTANAC >UAMH10320_GQ272617_SSU-LSU AGGGATCATTACCGAGTTCATGCCCCACAGCGGGGTAGATCTCCCACCCTTGTGTATTTATACCTGTGTTGCTTTGGCAGGCCGCTGGGCTGCGGTCTGGCCACCGGCTTCTTCGAAGCTGGTGCGTGCCTGCCAGAGGCCCCCCAAACTCTTGTTTGTCTAGTGTTGTCTGAGTATCATACCAATCGTTAAAACTTTCAACAACGGATCTCTTGGTTCGGGCATCGATGAAGAACGCAGCGAAATGCGATAAGTAATGCGAATTGCAGAATTCAGTGAATCATCGAATCTTTGAACGCACATTGCGCCCCTTGGTATTCCGAGGGGCATGCCTGTTCGAGCGTCATTTCAACCCCTCAAGCTCTGCTTGGTATTGGGCCTCGCCATCGCGGCGGGCCTTAAAATCAGTGGCGGTGCCGTCTCGGCTCCAAGCGTAGTAGCATCATCTCGCTCTGGAGACCCGGCGGTTGCTGGCCAGATAACCCCCAATTTTTTCTGTGGTTGACCTCGGATCAGGTAGGGATACCCGCTGAACTTAA |
IMAGES: