Collection Activities
Significant and interesting collections
To see what we have been up to in years past, please see our Annual Reports.
Fungus resistance testing and biodegradation fungi and bacteria. Many fungal isolates held at UAMH are listed as fungus resistance testing isolates or have been evaluated for biodegradation or bioremediation properties. Fungi expressing laccases and other ligninolytic enzymes, and having applications in metabolism of polycyclic aromatic hydrocarbons (PAH), dye decolorization or various industrial and environmental applications properties are listed here.
A bacterium Pseudomonas fluorescens strain LP6a (= UAMH 11620) has been extensively characterized by Dr. J. Foght (U of A Biological Sciences) and her colleagues and assessed for efficacy in bioremediation of oil spills. The bacterium was one of eleven isolates comprising the Standard Inocula used by Environment Canada to perform Oil Spill Bioremediation Agent efficacy tests (see publication list). Polycyclic aromatic hydrocarbons serve as carbon source for this microbe, but it also grows well on conventional organic substrates (e.g., glucose, acetate, Plate Count Agar). The organism grows at a temperature range of <4°C to 35°C. Pseudomonas fluorescens strain LP6a was isolated from soil contaminated by sulfur-rich petroleum condensate from a sour gas well blow-out in Lodgepole, Alberta in 1982. This link provides history for the incident: Lodepole blowout
Fungi associated with bats. North American bats are threatened by a disease called white nose syndrome caused by the fungus Geomyces destructans. Over four years, we cooperated with K. Vanderwolf, M. Sc. student from Univ of New Brunswick and NB Museum, who did extensive surveying of living hibernating bats in New Brunswick caves for incidence of the white nose pathogen G. destructans. In April 2011, the first cases of white nose syndrome and the presence of G. destructans was confirmed in New Brunswick bats. This devastating fungal disease continues to spread among bats in New Brunswick and in other parts of Canada and the US. In all, 260 UAMH accessions arose from Ms. Vanderwolf's study. These include large numbers of Geomyces species other than G. destructans as well as numerous rare or uncommon psychrophilic, keratinolytic, coprophilous and saprobic fungi of scientific interest (see Vanderwolf et al 2013a, 2013b). We verified or identified many of the accessioned isolates and are collaborating on taxonomic projects. Ms. Vanderwolf's thesis entitled "Characterizing the winter bat population, microclimate, and mycobiota of hiberating bats in New Brunswick caves" was completed in May 2012.
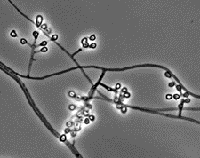
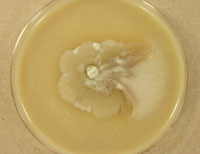
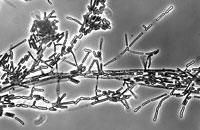
Geomyces species UAMH 11161 (left) and the mushroom Baeospora species UAMH 11492 (centre and right).
Fungi associated with tree bark beetles. The mountain pine beetle is responsible for destruction of large areas of lodgepole pine forests in western Canadian and is now moving into hybrid tree
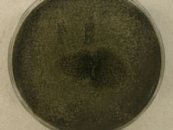 species as well as jackpine thereby increasing its potential for spread eastward in the boreal forest. Over 130 fungal isolates have been accessioned over several years through mountain pine beetle genomics projects. These isolates, representing six species of ophiostomatoid fungi (blue stain fungi), have been extensively genotyped (Tsui et al 2012; Tsui et al 2013). We continue to work with members of the UBC, U of A TRIA research group to voucher newly obtained mountain pine beetle associated fungi and to supply cultures for further biologic and genomic studies. Grosmannia clavigera isolates from different species of beetle and pine have been found to represent two phylogenetic species based on genetic and ecological assessment (see Alamouti et al Mol Ecol 2011). The isolates from these projects are listed in the UAMH catalogues but a complete list of recent acquisitions is available here.
species as well as jackpine thereby increasing its potential for spread eastward in the boreal forest. Over 130 fungal isolates have been accessioned over several years through mountain pine beetle genomics projects. These isolates, representing six species of ophiostomatoid fungi (blue stain fungi), have been extensively genotyped (Tsui et al 2012; Tsui et al 2013). We continue to work with members of the UBC, U of A TRIA research group to voucher newly obtained mountain pine beetle associated fungi and to supply cultures for further biologic and genomic studies. Grosmannia clavigera isolates from different species of beetle and pine have been found to represent two phylogenetic species based on genetic and ecological assessment (see Alamouti et al Mol Ecol 2011). The isolates from these projects are listed in the UAMH catalogues but a complete list of recent acquisitions is available here.
Ophiostomatoid fungi live in complex symbiotic relationships with the beetle and the availability of large numbers of well characterized isolates allows for study of the biological interactions between fungi, beetle and tree. The isolates represent different haplotypes and have been used for phylogenetic, population genetic and phylogenographic studies. Several molecular markers have been evaluated for their utility in identifying isolates among closely related blue stain fungi (see Roe et al Mol Ecol Res 2010; Roe et al Mol Ecol 2011). 
Genome sequencing. The genomes of two softwood ophiostomatoid fungi have been completed, including the pine pathogen Grosmannia clavigera (GenBank ACXQ02000000) for Grosmannia clavigera isolate SLKW1407 = UAMH 11150 (see DiGuistini et al 2011; DiGuistini et al 2009; Hesse-Orce et al 2010) and the pine saprophyte Ophiostoma piceae UAMH 11346 (GenBank AQHS00000000). Haridas et al. (2013) sequenced the genome for O. piceae and compared its transcriptomes with those of the mountain pine beetle associated pathogen Grosmannia clavigera UAMH 11150. They also demonstrated mating between two isolates of O. piceae UAMH 11346 and UAMH 11672. Ophiostoma piceae is a common saprophyte of pine and other softwood tree species.
Images show G. clavigera and O. piceae growing in culture.
Fungi associated with plant roots. The UAMH holds the largest North American repository of mycorrhizal and root associated fungi established from different hosts, habitats and regions in Canada and northern temperate regions. These vouchers are used by researchers to determine how endophytic fungi interact with their plant hosts, and their potential to improve plant growth particularly under conditions of environmental stress, but the ecological roles of many root endophytic fungi are still poorly understood.
Orchid fungi. The UAMH holds over 300 scientifically valuable isolates of orchid mycorrhizal fungi some of which have been shown to promote seed germination in vitro. Read more about conserving orchids and their fungal partners and our the role in the newly established North American Orchid Conservation Center.
Dark septate endophytes. In additional to mycorrhizal fungi, the UAMH houses numerous collections of plant root associated fungi for which the ecological function is less clear. Recently acquired collections include 34 strains of dark septate endophytes collectively known as members of the Phialocephala fortinii s.l. – Acephala applanata species complex (PAC).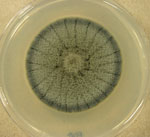
Acephala applanata UAMH 11715
Genome sequencing. The genomes of Meliniomyces bicolor strain E = UAMH 11274 obtained from Pinus sylvestris root tip in Scotland and Meliniomyces variabilis strain F (GT02V1) = UAMH 11265 have been fully sequenced. Meliniomyces was described by Hambleton and Sigler for isolates from ericoid and ectomycorrhizal roots that formed part of the Rhizoscyphus ericae aggregate.
Fungi accessioned in 2010 (and their associated sequences) included endophytes from Cenococcum geophilum mycorrhizae of Abies balsamea, Picea glauca or Betula papyrifera collected in Nova Scotia and Quebec (see Kernaghan 2011) and Meliniomyces species from ericoid and ectomycorrhizal hosts collected in forests of Scotland (see Grelet 2009, 2009, 2010).